When using an M-ary classifier on a binary classification problem, sometimes you want multiple dimensional output and sometimes you don’t. In the PRT the default is to supply only a single output but this can be specified by setting the twoClassParadigm property of classifiers.
Contents
A Quick Example
Consider a binary classification problem with an m-ary classifer (prtClassMap).
dsTrain = prtDataGenUnimodal; dsTest = prtDataGenUnimodal; classifier = prtClassMap; trainedClassifier = train(classifier, dsTrain); plot(trainedClassifier); title('Binary Classification with MAP','FontSize',16); output = run(trainedClassifier, dsTest); output.nFeatures
ans = 1
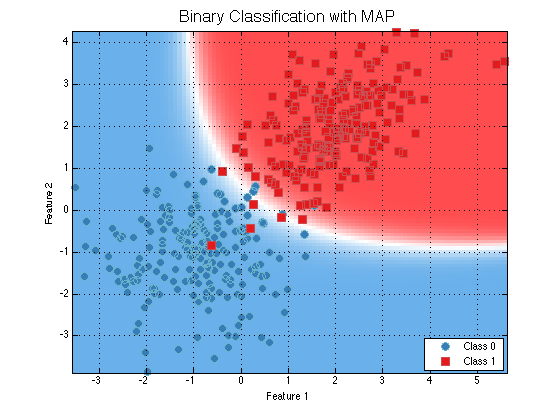
As you can see the output only has one feature. But consider the same M-ary classifier with a 3-class problem.
dsTrainMary = prtDataGenMary; dsTestMary = prtDataGenMary; trainedClassifierMary = train(classifier, dsTrainMary); plot(trainedClassifierMary); title('M-ary Classification with MAP','FontSize',16); outputMary = run(trainedClassifierMary, dsTestMary); outputMary.nFeatures
ans = 3
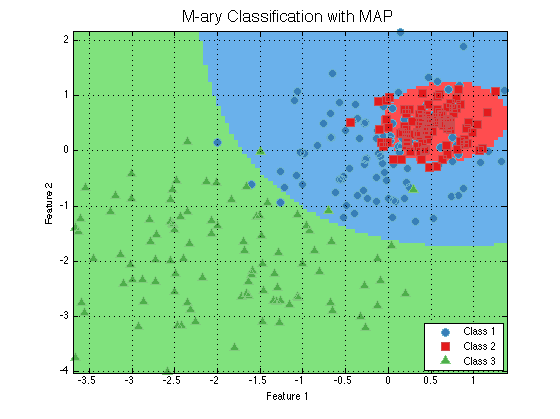
Now the output has 3 feautres. The trend continues with more classes, because prtClassMap declares itself as an Mary classififer
classifier.isNativeMary
ans = 1
the output is (as it should) have a column corresponding to the confidence of each class. Binary classification is an exception.
twoClassParadigm
By default the PRT checks when M-ary classifiers are run on binary data to see if it should output a single confidence or binary confidences. The mode of operation is stored in twoClassParadigm which by default is set to the string M-ary.
classifier.twoClassParadigm
ans = binary
If we return to the binary classification problem and set twoClassParadigm to mary we can see that we get the two outputs that we expect.
classifier.twoClassParadigm = 'mary';
trainedClassifierBinaryActingAsMary = train(classifier, dsTrain);
outputBinaryActingAsMary = run(trainedClassifierBinaryActingAsMary, dsTest);
outputBinaryActingAsMary.nFeatures
ans = 2
Why did you do that?
We debated, but ultimately we decided that it was less confusing this way. It is more rare to want both outputs for a binary classification problem than it is to expect a single. Most users expected this to run out of the box.
classifier = prtClassMap; trainedClassifier = train(classifier, dsTrain); output = run(trainedClassifier, dsTest); prtScoreRoc(output); title('Binary Classification with an M-ary Classifier','FontSize',16);
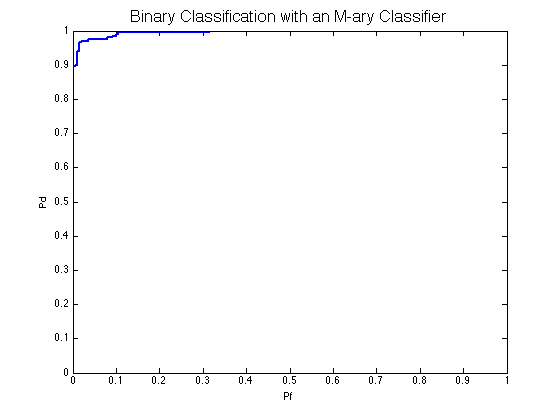
If you change the classifier to non-m-ary classification (prtClassGlrt for example) you expect the same code to work only by changing the classifier declaration.
classifier = prtClassGlrt; trainedClassifier = train(classifier, dsTrain); output = run(trainedClassifier, dsTest); prtScoreRoc(output); title('Binary Classification with a Binary Classifier','FontSize',16);
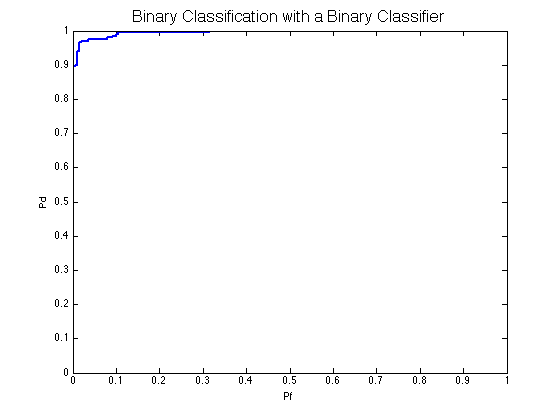
Without the twoClassParadigm system it wouldn’t be as easy to switch classifiers for binary problems as an extra step would be required to manually select which column of the output should be used to calculate the ROC.
Conclusion
Well that’s twoClassParadigm. It’s a convenient feature of the PRT that not many people know about because they don’t have to. In the rare cases when you want the confidence assigned to each class, you now know how to get them.